Analysis of Viral Promoter Usage in EBV-Infected Cell Lines: A Comparison of qPCR Following Conventional RNA Isolation and Nuclear Run-On Assay | SpringerLink
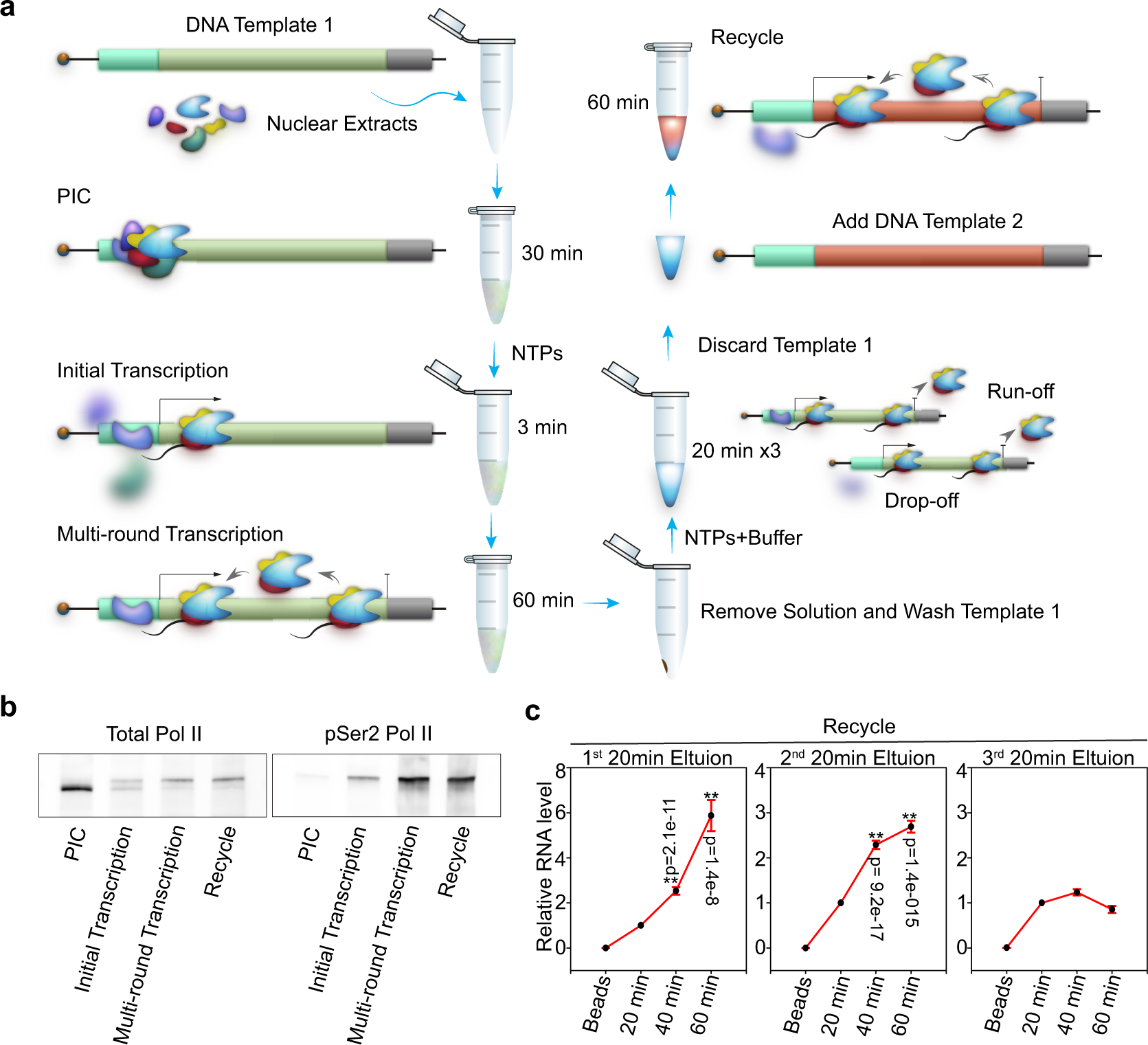
Transcription recycling assays identify PAF1 as a driver for RNA Pol II recycling | Nature Communications

Analysis of Termination of Transcription Using BrUTP-strand-specific Transcription Run-on (TRO) Approach
Time-Dependent c-Myc Transactomes Mapped by Array-Based Nuclear Run-On Reveal Transcriptional Modules in Human B Cells | PLOS ONE

Comparison of the conventional run-off transcription and the ribozyme... | Download Scientific Diagram
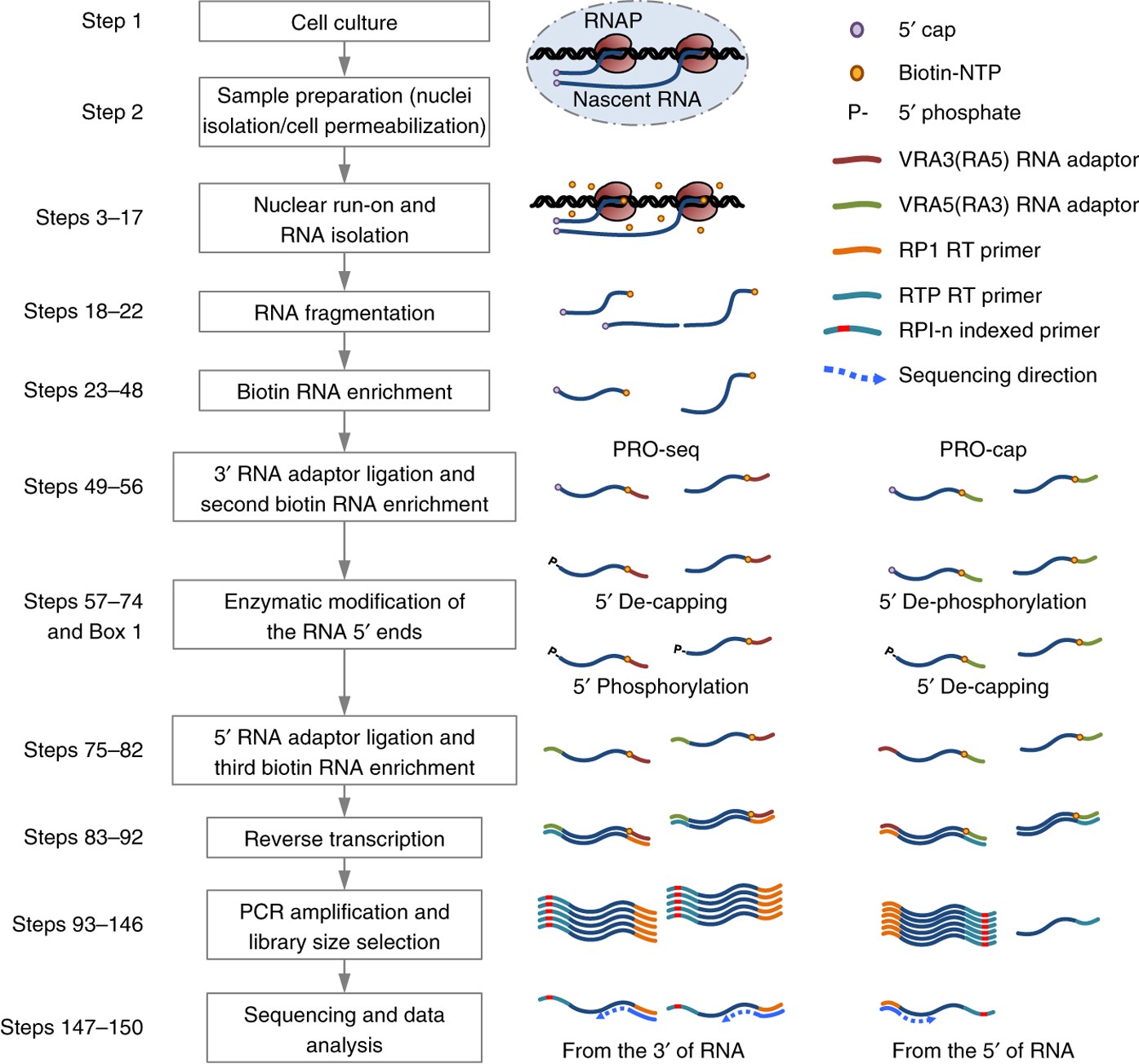
Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq) | Nature Protocols
The Basics of Transcription - I Transcription Regulation And Gene Expression in Eukaryotes FS 2014 Graduate Course G2